
Security News
Research
Data Theft Repackaged: A Case Study in Malicious Wrapper Packages on npm
The Socket Research Team breaks down a malicious wrapper package that uses obfuscation to harvest credentials and exfiltrate sensitive data.
This library contains the trained graph neural network model for the prediction of homolytic bond dissociation energies (BDEs) of organic molecules with C, H, N, and O atoms. This package offers a command-line interface to the web-based model predictions at bde.ml.nrel.gov.
The basic interface works as follows, where predict expects a list of SMILES strings of the target molecules
>>> from alfabet import model
>>> model.predict(['CC', 'NCCO'])
molecule bond_index bond_type fragment1 fragment2 ... bde_pred is_valid
0 CC 0 C-C [CH3] [CH3] ... 90.278282 True
1 CC 1 C-H [H] [CH2]C ... 99.346184 True
2 NCCO 0 C-N [CH2]CO [NH2] ... 89.988495 True
3 NCCO 1 C-C [CH2]O [CH2]N ... 82.122429 True
4 NCCO 2 C-O [CH2]CN [OH] ... 98.250961 True
5 NCCO 3 H-N [H] [NH]CCO ... 99.134750 True
6 NCCO 5 C-H [H] N[CH]CO ... 92.216087 True
7 NCCO 7 C-H [H] NC[CH]O ... 92.562988 True
8 NCCO 9 H-O [H] NCC[O] ... 105.120598 True
The model breaks all single, non-cyclic bonds in the input molecules and calculates their bond dissociation energies. Typical prediction errors are less than 1 kcal/mol. The model is based on Tensorflow (2.x), and makes heavy use of the neural fingerprint library (0.1.x).
For additional details, see the publication: St. John, P. C., Guan, Y., Kim, Y., Kim, S., & Paton, R. S. (2020). Prediction of organic homolytic bond dissociation enthalpies at near chemical accuracy with sub-second computational cost. Nature Communications, 11(1). doi:10.1038/s41467-020-16201-z
Note: For the exact model described in the text, install alfabet version 0.0.x. Versions >0.1 have been updated for tensorflow 2.
Installation with conda is recommended, as rdkit can otherwise be difficult to install
$ conda create -n alfabet -c conda-forge python=3.7 rdkit
$ source activate alfabet
$ pip install alfabet
``
FAQs
A library to estimate bond dissociation energies (BDEs) of organic molecules
We found that alfabet demonstrated a healthy version release cadence and project activity because the last version was released less than a year ago. It has 1 open source maintainer collaborating on the project.
Did you know?

Socket for GitHub automatically highlights issues in each pull request and monitors the health of all your open source dependencies. Discover the contents of your packages and block harmful activity before you install or update your dependencies.

Security News
Research
The Socket Research Team breaks down a malicious wrapper package that uses obfuscation to harvest credentials and exfiltrate sensitive data.
Research
Security News
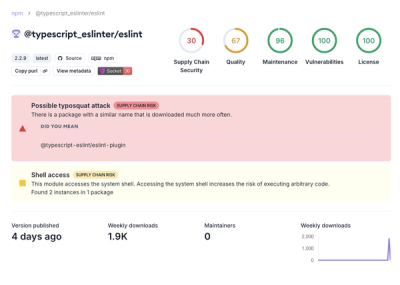
Attackers used a malicious npm package typosquatting a popular ESLint plugin to steal sensitive data, execute commands, and exploit developer systems.

Security News
The Ultralytics' PyPI Package was compromised four times in one weekend through GitHub Actions cache poisoning and failure to rotate previously compromised API tokens.