
Security News
Research
Data Theft Repackaged: A Case Study in Malicious Wrapper Packages on npm
The Socket Research Team breaks down a malicious wrapper package that uses obfuscation to harvest credentials and exfiltrate sensitive data.
Med-Imagetools: Transparent and Reproducible Medical Image Processing Pipelines in Python
nnunet nnU-Net compatibility moderandom_state for reproducible seedscontinue_processing flag to continue autopipelinedry_run flag to only crawl the datasetMed-Imagetools, a python package offers the perfect tool to transform messy medical dataset folders to deep learning ready format in few lines of code. It not only processes DICOMs consisting of different modalities (like CT, PET, RTDOSE and RTSTRUCTS), it also transforms them into deep learning ready subject based format taking the dependencies of these modalities into consideration.
A medical dataset, typically contains multiple different types of scans for a single patient in a single study. As seen in the figure below, the different scans containing DICOM of different modalities are interdependent on each other. For making effective machine learning models, one ought to take different modalities into account.

Fig.1 - Different network topology for different studies of different patients
Med-Imagetools is a unique tool, which focuses on subject based Machine learning. It crawls the dataset and makes a network by connecting different modalities present in the dataset. Based on the user defined modalities, med-imagetools, queries the graph and process the queried raw DICOMS. The processed DICOMS are saved as nrrds, which med-imagetools converts to torchio subject dataset and eventually torch dataloader for ML pipeline.

Fig.2 - Med-Imagetools AutoPipeline diagram
pip install med-imagetools
conda create -n mit
conda activate mit
pip install med-imagetools
conda create -n mit
conda activate mit
pip install -e git+https://github.com/bhklab/med-imagetools.git
This will install the package in editable mode, so that the installed package will update when the code is changed.
imgtools
[!WARNING] Warning: This feature is still in beta. Use with caution and report any issues on GitHub.

torch and torchio dependencies have been moved to an extra pip install flag. Use pip install med-imagetools[torch].Med-Imagetools takes two step approch to turn messy medical raw dataset to ML ready dataset.
Autopipeline: Crawls the raw dataset, forms a network and performs graph query, based on the user defined modalities. The relevant DICOMS, get processed and saved as nrrds
autopipeline\
[INPUT DIRECTORY] \
[OUTPUT DIRECTORY] \
--modalities [str: CT,RTSTRUCT,PT] \
--spacing [Tuple: (int,int,int)]\
--n_jobs [int]\
--visualize [flag]\
--nnunet [flag]\
--train_size [float]\
--random_state [int]\
--roi_yaml_path [str]\
--continue_processing [flag]\
--dry_run [flag]
class Dataset: This class converts processed nrrds to torchio subjects, which can be easily converted to torch dataset
from imgtools.io import Dataset
subjects = Dataset.load_from_nrrd(output_directory, ignore_multi=True)
data_set = tio.SubjectsDataset(subjects)
data_loader = torch.utils.data.DataLoader(data_set, batch_size=4, shuffle=True, num_workers=4)
These google collab notebooks will introduce the main functionalities of med-imagetools. More information can be found here
Thanks to the following people who have contributed to this project:
This project uses the following license: MIT License
FAQs
Med-Imagetools: Transparent and Reproducible Medical Image Processing Pipelines in Python
We found that med-imagetools demonstrated a healthy version release cadence and project activity because the last version was released less than a year ago. It has 2 open source maintainers collaborating on the project.
Did you know?

Socket for GitHub automatically highlights issues in each pull request and monitors the health of all your open source dependencies. Discover the contents of your packages and block harmful activity before you install or update your dependencies.

Security News
Research
The Socket Research Team breaks down a malicious wrapper package that uses obfuscation to harvest credentials and exfiltrate sensitive data.
Research
Security News
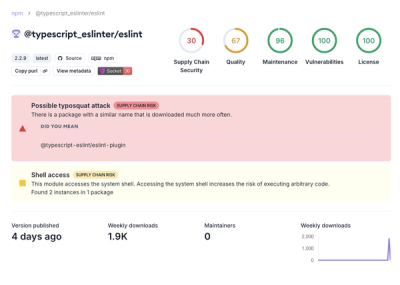
Attackers used a malicious npm package typosquatting a popular ESLint plugin to steal sensitive data, execute commands, and exploit developer systems.

Security News
The Ultralytics' PyPI Package was compromised four times in one weekend through GitHub Actions cache poisoning and failure to rotate previously compromised API tokens.