Research
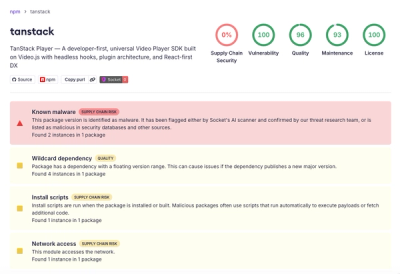
Malicious npm Package Brand-Squats TanStack to Exfiltrate Environment Variables
A brand-squatted TanStack npm package used postinstall scripts to steal .env files and exfiltrate developer secrets to an attacker-controlled endpoint.
pydustmasker
Advanced tools
Python bindings to DustMasker, a utility to identify and mask low-complexity regions in nucleotide sequences
pydustmasker is a Python library that enables efficient detection and masking of low-complexity regions in nucleotide sequences using the SDUST1 and Longdust2 algorithms.
The full documentation for pydustmasker, including installation instructions, theoretical background, and API reference, is available at https://apcamargo.github.io/pydustmasker.
Using pip:
pip install pydustmasker
Using pixi:
# Create a new Pixi workspace and navigate into the workspace directory
pixi init my_workspace && cd my_workspace
# Add Bioconda to the list of channels of your Pixi workspace
pixi workspace channel add bioconda
# Add pydustmasker to your Pixi workspace
pixi add pydustmasker
To identify and mask low-complexity regions in a nucleotide sequence, create an instance of a masker class and provide your sequence to it. A masker class implements a specific low-complexity detection algorithm and provides methods to retrieve the detected regions and to generate a masked version of the sequence. pydustmasker provides two such classes, corresponding to different detection algorithms: SDUST and Longdust. The SDUST algorithm is implemented in the DustMasker class, while the Longdust algorithm is implemented in the LongdustMasker class.
>>> import pydustmasker
# Example nucleotide sequence
>>> seq = "CGTATATATATAGTATGCGTACTGGGGGGGCT"
# Create a DustMasker object to identify low-complexity regions with the SDUST algorithm
>>> masker = pydustmasker.DustMasker(seq)
# The len() function returns the number of low-complexity regions detected in the sequence
>>> len(masker)
1
# Get the number of bases within low-complexity regions and the intervals of these regions
>>> masker.n_masked_bases
7
>>> masker.intervals
((23, 30),)
# The masker object is iterable, yielding start and end positions of each low-complexity region
>>> for start, end in masker: # (4)!
... print(f"{start}-{end}: {seq[start:end]}")
23-30: GGGGGGG
You can generate a masked version of the sequence using the mask() method. By default, low-complexity regions are soft-masked by converting bases to lowercase. Setting the hard parameter to True enables hard-masking, in which affected bases are replaced with the ambiguous nucleotide N.
# The mask() method returns the sequence with low-complexity regions soft-masked
>>> masker.mask()
'CGTATATATATAGTATGCGTACTgggggggCT'
# Hard-masking can be enabled by setting the `hard` parameter to `True`
>>> masker.mask(hard=True)
'CGTATATATATAGTATGCGTACTNNNNNNNCT'
The identification of low-complexity regions can be tuned via algorithm-specific parameters. Both DustMasker and LongdustMasker provide multiple options, documented in the API reference, that control how low-complexity regions are determined. One shared parameter is score_threshold, which controls detection stringency: lowering this threshold results in more regions being classified as low-complexity, whereas increasing it restricts detection to the most clearly low-complexity regions.
# Setting `score_threshold` to 10 results in more low-complexity regions being detected
>>> masker = pydustmasker.DustMasker(seq, score_threshold=10)
>>> len(masker)
2
>>> masker.intervals
((2, 12), (23, 30))
>>> masker.mask()
'CGtatatatataGTATGCGTACTgggggggCT'
When working with large numbers of sequences, you can run pydustmasker in parallel to process multiple sequences at the same time. This can substantially reduce the total time needed to process all sequences.
The example below uses Biopython to parse a FASTA file containing multiple sequences, which are then processed in parallel using a pool of worker processes from the multiprocessing module. Each sequence record is submitted to the worker pool via imap and processed with LongdustMasker to identify low-complexity regions using the Longdust algorithm. The resulting intervals are written to the output file as they become available.
#!/usr/bin/env python
import multiprocessing.pool
from Bio import SeqIO
import pydustmasker
input_file = "sequences.fna"
output_file = "lc_intervals.tsv"
def process_record(record):
masker = pydustmasker.LongdustMasker(str(record.seq), score_threshold=12)
return record.id, masker.intervals
if __name__ == "__main__":
with open(output_file, "w") as f, multiprocessing.pool.Pool() as pool:
records = SeqIO.parse(input_file, "fasta")
for name, intervals in pool.imap(process_record, records):
for start, end in intervals:
f.write(f"{name}\t{start}\t{end}\n")
Morgulis, Aleksandr, et al. A Fast and Symmetric DUST Implementation to Mask Low-Complexity DNA Sequences. Journal of Computational Biology, vol. 13, no. 5, June 2006, pp. 1028–40. https://doi.org/10.1089/cmb.2006.13.1028. ↩
Li, Heng, and Brian Li. Finding Low-Complexity DNA Sequences with Longdust. arXiv, 2025. https://doi.org/10.48550/arxiv.2509.07357. ↩
FAQs
Python bindings to DustMasker, a utility to identify and mask low-complexity regions in nucleotide sequences
We found that pydustmasker demonstrated a healthy version release cadence and project activity because the last version was released less than a year ago. It has 1 open source maintainer collaborating on the project.
Did you know?

Socket for GitHub automatically highlights issues in each pull request and monitors the health of all your open source dependencies. Discover the contents of your packages and block harmful activity before you install or update your dependencies.

Research
A brand-squatted TanStack npm package used postinstall scripts to steal .env files and exfiltrate developer secrets to an attacker-controlled endpoint.
Research
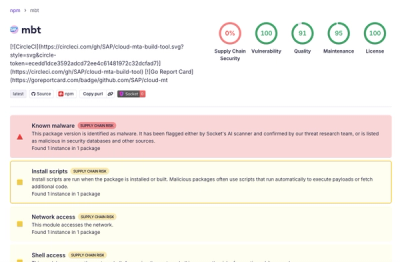
Compromised SAP CAP npm packages download and execute unverified binaries, creating urgent supply chain risk for affected developers and CI/CD environments.

Company News
Socket has acquired Secure Annex to expand extension security across browsers, IDEs, and AI tools.