
Security News
High Salaries No Longer Enough to Attract Top Cybersecurity Talent
A survey of 500 cybersecurity pros reveals high pay isn't enough—lack of growth and flexibility is driving attrition and risking organizational security.
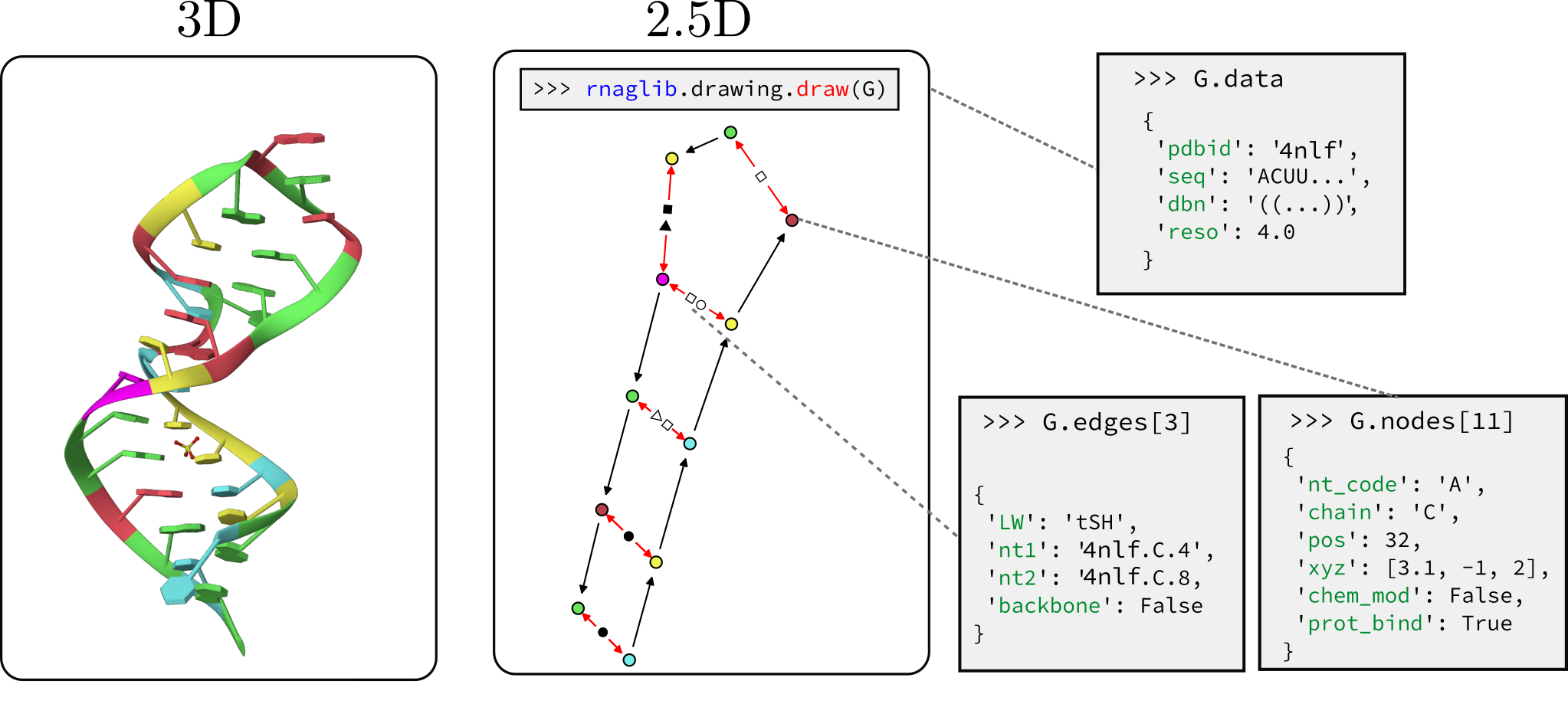
RNAglib: Tools for learning on the structure of RNA using 2.5D geometric representations

rnaglib)RNAglib is a Python package for studying RNA 2.5D and 3D structures. Functionality includes automated data loading,
analysis,
visualization, ML model building and benchmarking.
We host RNAs annotated with molecule, base pair, and nucleotide level attributes. These include, but are not limited to:
The package can be cloned and the source code used directly. We also deploy it as a pip package and recommend using this install in conda environments.
If one wants to use GPU support, one should install Pytorch and DGL with the appropriate options. Otherwise, you can just skip this step and the pip installs of Pytorch and DGL will be used.
Then, one just needs to run :
pip install rnaglib
Then one can start using the packages functionalities by importing them in one's python script.
Optional Dependencies
We use fr3d-python to build 2.5D annotations from PDBs and mmCIFs. This has to be installed manually with:
pip install git+https://github.com/cgoliver/fr3d-python.git
To load graphs and train with dgl:
pip install dgl
To load graphs and train with pytorch geometric:
pip install torch_geometric
Advanced data splitting of datasets (i.e. by sequence or structure-based similarity) depends on executables cd-hit and RNAalign. You may install these yourself or use the convenience script provided in this repo as follows, though we do not guarantee it will work on any system and has only bee tested on linux:
chmod u+x install_dependencies.sh
./install_dependencies.sh /my/path/to/executables
Run the rnaglib_download script to fetch the database of annotated structures. By default it will download to ~/.rnaglib.
Use the --help flag for more options.
$ rnaglib_download
In addition to analysing RNA data, RNAglib also distributes available parsed RNA structures.
Databases of annotated structures can be downloaded directly from Zenodo.
They can also be obtained through the provided command line utility $ rnaglib_download -r all|nr.
| Version | Date | Total RNAs | Total Non-Redundant | Non-redundant version | rnaglib commit |
|---|---|---|---|---|---|
| 2.0.2 | 25-02-25 | 8441 | 2921 | 3.375 | ac303c7 |
| 2.0.0 | 12-01-25 | 8305 | 2877 | 3.369 | 33a9e989 |
| 1.0.0 | 15-02-23 | 5759 | 1176 | 3.269 | 5446ae2c |
| 0.0.0 | 20-07-21 | 3739 | 899 | 3.186 | eb25dabd |
To speed up some functions we also build an efficient index of the data. This is needed for the rnaglib.tasks functionality.
$ rnaglib_index
rnaglib?Datasets of RNA 3D structures ready-to-use for machine learning model benchmarking in various biologically relevant tasks.
All tasks take as input an RNA 3D structure and have as output either residue or whole RNA-level properties.
We currently support the following 6 prediction tasks and the ability to create new tasks:
See docs for more info on usage.
Each link for the tasks above takes you to a demo.py file with example usage for each task.
Everything you need to train and evaluate a model is built on 3 basic ingredients:
rnaglib.Task object with holds all the relevant data, splits and
functionality.rnaglib.Representation object which converts raw RNAs to tensor
formats.rnaglib.learning.PyGmodelfrom rnaglib.tasks import ChemicalModification
from rnaglib.transforms import GraphRepresentation
from rnaglib.learning.task_models import PygModel
# Load task, representation, and get loaders
task = ChemicalModification(root="my_root")
model = PygModel.from_task(task)
pyg_rep = GraphRepresentation(framework="pyg")
task.add_representation(pyg_rep)
train_loader, val_loader, test_loader = task.get_split_loaders(batch_size=8)
for batch in train_loader:
batch = batch['graph'].to(model.device)
output = model(batch)
test_metrics = model.evaluate(task, split='test')
By default, most tasks use the residue type as the only residue-level feature and if you choose a graph representation, the graph is computed using the Leontis Westhof nomenclature.
Features are handled through the rnaglib.Transforms class. Each
Transform is a callable which receives an RNA, applies any operation to it
(e.g. adding a piece of data) and returns it.
We provide some Transforms which you can use to add more features to datasets
by simply passing it to the Task.add_feature() method.
This is an example of adding RNAFM embeddings as features to a dataset.
from rnaglib.tasks import RNAGo
from rnaglib.transforms import RNAFMTransform
# Take out the ingredients
task = RNAGo(root="go")
tr = RNAFMTransform()
pyg_rep = GraphRepresentation(framework="pyg")
model = PygModel.from_task(task, num_node_features=644)
task.add_feature(tr)
task.add_representation(pyg_rep)
for batch in train_loader:
batch = batch['graph'].to(model.device)
output = model(batch)
test_metrics = model.evaluate(task, split='test')
Since the RNAFM embeddings have 640 dimensions, and the nucleotide tensor is 4 dimensional, we initialize the model with 644 dimensions.
Current release contains annotations generated by x3dna-dssr as well as some additional ones that we added for all available PDBs at the time of release.
Each RNA is stored as a networkx graph where nodes are residues and edges are backbone and base pairing edges. The networkx graph object has graph-level, node-level and edge-level attributes. Here is a reference for all the annotations currently available.
>>> from rnaglib.data_loading import rna_from_pdbid
>>> rna_dict = graph_from_pdbid('1fmn') # fetch from local database or RCSB if not found
>>> rna_dict['rna'].graph
{'name': '1fmn', 'pdbid': '1fmn', 'ligands': [{'id': ('H_FMN', 36, ' '), 'name': 'FMN', 'smiles': 'Cc1cc2c(cc1C)N(C3=NC(=O)NC(=O)C3=N2)CC(C(C(COP(=O)(O)O)O)O)O', 'rna_neighs': ['1fmn.A.10', '1fmn.A.11', '1fmn.A.12', '1fmn.A.13', '1fmn.A.24', '1fmn.A.25', '1fmn.A.26', '1fmn.A.27', '1fmn.A.28', '1fmn.A.7', '1fmn.A.8', '1fmn.A.9']}], 'ions': []}
You can extract Leontis-Westhof interactions and convert 3D structures to 2.5D graphs. We wrap a fork of fr3d-python to support this functionality.
from rnaglib.prepare_data import fr3d_to_graph
G = fr3d_to_graph("../data/structures/1fmn.cif")
Warning: this method currently does not support non-standard residues. Support coming soon. Up to version 1.0.0 of the RNA database were created using x3dna-dssr which do contain non-standard residues.
We customize networkx graph drawing functionalities to give some convenient visualization of 2.5D base pairing networks. This is not a dedicated visualization tool, it is only intended for quick debugging. We point you to VARNAhttps://varna.lisn.upsaclay.fr/ or RNAscape for a full-featured visualizer.
from rnaglib.drawing import rna_draw
rna_draw(G, show=True, layout="spring")
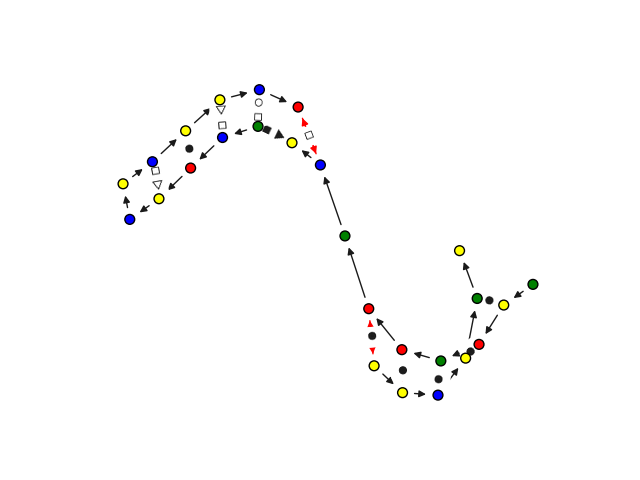
When dealing with 3D structures as 2.5D graphs we support graph-level comparison through the graph edit distance.
from rnaglib.ged import graph_edit_distance
from rnaglib.utils import graph_from_pdbid
G = graph_from_pdbid("4nlf")
print(graph_edit_distance(G, G)) # 0.0
Go to root of the rnaglib directory from a git clone and run pytest.
pip install pytest
pytest tests/
@article{mallet2022rnaglib,
title={RNAglib: a python package for RNA 2.5 D graphs},
author={Mallet, Vincent and Oliver, Carlos and Broadbent, Jonathan and Hamilton, William L and Waldisp{\"u}hl, J{\'e}r{\^o}me},
journal={Bioinformatics},
volume={38},
number={5},
pages={1458--1459},
year={2022},
publisher={Oxford University Press}
}
rnaglibIf you use rnaglib in one of your projects, please cite and feel free to make a pull request so we can list your project here.
rnaglib@cs.mcgill.caFAQs
RNAglib: Tools for learning on the structure of RNA using 2.5D geometric representations
We found that rnaglib demonstrated a healthy version release cadence and project activity because the last version was released less than a year ago. It has 2 open source maintainers collaborating on the project.
Did you know?

Socket for GitHub automatically highlights issues in each pull request and monitors the health of all your open source dependencies. Discover the contents of your packages and block harmful activity before you install or update your dependencies.

Security News
A survey of 500 cybersecurity pros reveals high pay isn't enough—lack of growth and flexibility is driving attrition and risking organizational security.

Product
Socket, the leader in open source security, is now available on Google Cloud Marketplace for simplified procurement and enhanced protection against supply chain attacks.

Security News
Corepack will be phased out from future Node.js releases following a TSC vote.