Product
Announcing Socket Fix 2.0
Socket Fix 2.0 brings targeted CVE remediation, smarter upgrade planning, and broader ecosystem support to help developers get to zero alerts.
A Python package for directed evolution on a protein sequence with gradient-based discrete Markov chain monte carlo (MCMC). Users are able to compose custom models that map sequence to function with pretrained models, including protein language models (PLMs), to guide and constrain search. Our package natively integrates with 🤗 HuggingFace and supports PLMs from transformers.
Our MCMC sampler identifies promising amino acids to mutate via model gradients taken with respect to the input (i.e., sensitivity analysis). We allow users to compose their own custom target function for MCMC by leveraging the Product of Experts MCMC paradigm. Each model is an "expert" that contributes its own knowledge about the protein's fitness landscape to the overall target function. The sampler is designed to be more efficient and effective than brute force and random search while maintaining most of the generality and flexibility.
See our publication and our documentation for more details.
EvoProtGrad is available on PyPI and can be installed with pip:
pip install evo_prot_grad
For the bleeding edge version, and/or if you wish to run tests or register a new expert model with EvoProtGrad, please clone this repo and install in editable mode as follows:
git clone https://github.com/NREL/EvoProtGrad.git
cd EvoProtGrad
pip install -e .
Test the code by running python3 -m unittest.
See demo.ipynb to get started right away in a Jupyter notebook or
Create a ProtBERT expert from a pretrained HuggingFace protein language model (PLM) using evo_prot_grad.get_expert:
import evo_prot_grad
prot_bert_expert = evo_prot_grad.get_expert('bert', scoring_strategy = 'pseudolikelihood_ratio', temperature = 1.0)
The default BERT-style PLM in EvoProtGrad is Rostlab/prot_bert. Normally, we would need to also specify the model and tokenizer. When using a default PLM expert, we automatically pull these from the HuggingFace Hub. The temperature parameter rescales the expert scores and can be used to trade off the importance of different experts. The pseudolikelihood_ratio strategy computes the ratio of the "pseudo" log-likelihood (this isn't the exact log-likelihood when the protein language model is a masked language model) of the wild type and mutant sequence.
Then, create an instance of DirectedEvolution and run the search, returning a list of the best variant per Markov chain (as measured by the prot_bert expert):
variants, scores = evo_prot_grad.DirectedEvolution(
wt_fasta = 'test/gfp.fasta', # path to wild type fasta file
output = 'best', # return best, last, all variants
experts = [prot_bert_expert], # list of experts to compose
parallel_chains = 1, # number of parallel chains to run
n_steps = 20, # number of MCMC steps per chain
max_mutations = 10, # maximum number of mutations per variant
verbose = True # print debug info to command line
)()
We provide a few experts in evo_prot_grad/experts that you can use out of the box, such as:
Protein Language Models (PLMs)
bert, BERT-style PLMs, default: Rostlab/prot_bertcausallm, CausalLM-style PLMs, default: lightonai/RITA_sesm, ESM-style PLMs, default: facebook/esm2_t6_8M_UR50DPotts models
evcouplingsand an generic expert for supervised downstream regression models
onehot_downstream_regressionIf you use EvoProtGrad in your research, please cite the following publication:
@article{emami2023plug,
title={Plug \& play directed evolution of proteins with gradient-based discrete MCMC},
author={Emami, Patrick and Perreault, Aidan and Law, Jeffrey and Biagioni, David and John, Peter St},
journal={Machine Learning: Science and Technology},
volume={4},
number={2},
pages={025014},
year={2023},
publisher={IOP Publishing}
}
FAQs
Directed evolution of proteins with fast gradient-based discrete MCMC.
We found that evo-prot-grad demonstrated a healthy version release cadence and project activity because the last version was released less than a year ago. It has 1 open source maintainer collaborating on the project.
Did you know?

Socket for GitHub automatically highlights issues in each pull request and monitors the health of all your open source dependencies. Discover the contents of your packages and block harmful activity before you install or update your dependencies.

Product
Socket Fix 2.0 brings targeted CVE remediation, smarter upgrade planning, and broader ecosystem support to help developers get to zero alerts.

Security News
Socket CEO Feross Aboukhadijeh joins Risky Business Weekly to unpack recent npm phishing attacks, their limited impact, and the risks if attackers get smarter.
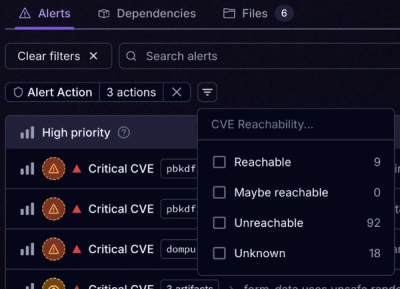
Product
Socket’s new Tier 1 Reachability filters out up to 80% of irrelevant CVEs, so security teams can focus on the vulnerabilities that matter.