Product
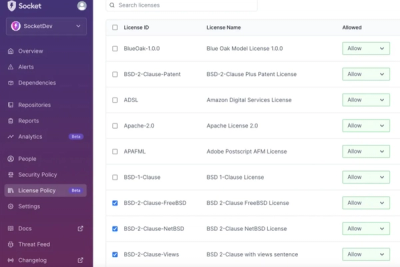
Introducing License Enforcement in Socket
Ensure open-source compliance with Socket’s License Enforcement Beta. Set up your License Policy and secure your software!
Tools for phylogenetic data analysis including visualization and cluster-computing support.
.. image:: https://img.shields.io/pypi/v/phylotoast.svg?style=plastic
:target: https://pypi.python.org/pypi/phylotoast
:alt: Latest Version
.. image:: https://img.shields.io/pypi/l/phylotoast.svg?style=plastic
:target: https://pypi.python.org/pypi/phylotoast
:alt: License
.. image:: https://img.shields.io/pypi/format/phylotoast.svg?style=plastic
:target: https://pypi.python.org/pypi/phylotoast
:alt: Download format
.. image:: https://img.shields.io/travis/smdabdoub/phylotoast.svg?style=plastic
:target: https://travis-ci.org/smdabdoub/phylotoast
:alt: Travis CI build status
.. image:: https://img.shields.io/badge/install%20with-bioconda-4682b4.svg?style=plastic
:target: https://bioconda.github.io/recipes/phylotoast/README.html
The PhyloToAST project is a collection of python code and scripts that modify the QIIME [1] pipeline by adding/changing several steps including: support for cluster-computing, multiple primer support (eliminate primer bias) [2], enhanced support for species-specific analysis, and additional visualization tools.
To install PhyloToAST from PyPI:
.. code-block:: bash
$ pip install phylotoast
From source:
.. code-block:: bash
$ python setup.py install
Full documentation for the scripts and code is available at
docs.phylotoast.org_ (hosted by Read the Docs_)
The list of required modules will vary depending on which executable scripts and/or parts of the API you may use. For this reason there are no required dependencies that will be automatically installed along with PhyloToAST. Each executable script will check that the required libraries are installed and will print a message if any are not found.
If you would like to install everything up front, the following is a complete list of libraries that are used in PhyloToAST:
numpy_scipy_matplotlib_ >= 1.5.0biopython_ >= 1.60scikit-bio_scikit-learn_pandas_statsmodels_palettable_biom-format_ >= 2.1.5h5py_ (for parsing BIOM v2.x format files)The PhyloToAST source_ is hosted on github.
Dabdoub, S. M. et al. PhyloToAST: Bioinformatics tools for species-level analysis and visualization of complex microbial datasets. Sci. Rep. 6, 29123; doi: 10.1038/srep29123 (2016).
Tsigarida and Dabdoub et al., The Influence of Smoking on the Peri-Implant
Microbiome. Journal of Dental Research, 2015; doi: 10.1177/0022034515590581_
Mason et al., The subgingival microbiome of clinically healthy current
and never smokers. The ISME Journal, 2014; doi:10.1038/ismej.2014.114_
Dabdoub et al., Patient-specific Analysis of Periodontal and Peri-implant Microbiomes.
Journal of Dental Research, 2013; doi: 10.1177/0022034513504950_
[1] J Gregory Caporaso, et al., QIIME allows analysis of
high-throughput community sequencing data. Nature Methods, 2010;
doi:10.1038/nmeth.f.303_
[2] Kumar PS, et al., Target Region Selection Is a Critical Determinant
of Community Fingerprints Generated by 16S Pyrosequencing. PLoS ONE
(2011) 6(6): e20956. doi:10.1371/journal.pone.0020956_
.. _docs.phylotoast.org: http://docs.phylotoast.org
.. _Read the Docs: http://readthedocs.org
.. _numpy: http://numpy.org
.. _scipy: http://scipy.org
.. _matplotlib: http://matplotlib.org
.. _biopython: http://biopython.org
.. _scikit-bio: http://scikit-bio.org
.. _scikit-learn: http://scikit-learn.org
.. _pandas: http://pandas.pydata.org
.. _statsmodels: http://statsmodels.sourceforge.net/
.. _palettable: https://jiffyclub.github.io/palettable/
.. _biom-format: http://biom-format.org
.. _h5py: http://www.h5py.org/
.. _PhyloToAST source: http://github.com/smdabdoub/phylotoast
.. _doi: 10.1177/0022034515590581: http://dx.doi.org/10.1177/0022034515590581
.. _doi:10.1038/ismej.2014.114: http://dx.doi.org/10.1038/ismej.2014.114
.. _doi: 10.1177/0022034513504950: http://dx.doi.org/10.1177/0022034513504950
.. _doi:10.1038/nmeth.f.303: http://dx.doi.org/10.1038/nmeth.f.303
.. _doi:10.1371/journal.pone.0020956: http://dx.doi.org/10.1371/journal.pone.0020956
FAQs
Tools for phylogenetic data analysis including visualization and cluster-computing support.
We found that phylotoast demonstrated a healthy version release cadence and project activity because the last version was released less than a year ago. It has 1 open source maintainer collaborating on the project.
Did you know?

Socket for GitHub automatically highlights issues in each pull request and monitors the health of all your open source dependencies. Discover the contents of your packages and block harmful activity before you install or update your dependencies.

Product
Ensure open-source compliance with Socket’s License Enforcement Beta. Set up your License Policy and secure your software!
Product
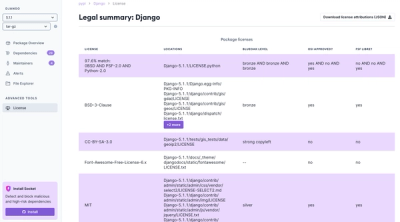
We're launching a new set of license analysis and compliance features for analyzing, managing, and complying with licenses across a range of supported languages and ecosystems.
Product
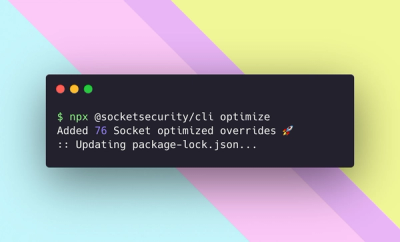
We're excited to introduce Socket Optimize, a powerful CLI command to secure open source dependencies with tested, optimized package overrides.