Product
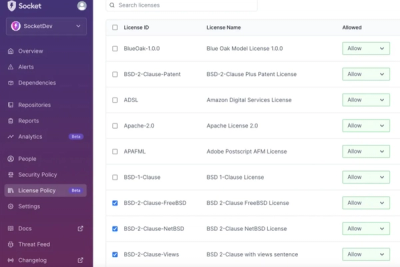
Introducing License Enforcement in Socket
Ensure open-source compliance with Socket’s License Enforcement Beta. Set up your License Policy and secure your software!
pyroscopegriddingcpu
Advanced tools
Data fusion package for transforming L2 satellite to L3 spatial-temporal gridded data
Author: Sally Zhao, Neil Gutkin
It is important to monitor Earth system data for both research and scientific reasons — analysis of such data furthers understanding of the planet and better informs political, economic, and policy decisions. Thus, in an effort to aid the Making Earth System Data Records for Use in Research Environments (MEaSUREs) Program, this project developed Python code for fusing six satellite Level 2 aerosol data (three are from geostationary satellites (GEO), and other three are from low earth orbital satellites (LEO)) from Dark Target Aerosol Retrieval Algorithm. This work emulated existing IDL workflow, which was originally created with IDL (interactive data language) by the science team. Quality checking was done by comparison to Panoply results and netCDF output matching through Python. Fused datasets generated by the program allow for visualization and analysis of the global aerosol data record over specific time periods. It also aids in research and analysis as users can better manipulate and work with satellite and sensor data. By making such code, and the accompanying functionality, open source and scalable, the scientific community is granted easier access to aerosol data processing resources.
Required libraries:
check requirements.txt for the required versions or additional libraries. numba is not required for the cpu-only package.
Supported input formats include netCDF4 and HDF file formats.
Depending on export selection, the package can create netCDF4 or geoTIFF files.
Users can manually call functions from the command line. User specifications are demarked by flags.
-fn (filename): Single filename to read in.
-fl (filelist): Location of file that contains a list of files (and their locations) to read in.
-gl (geolocation variables): Geolocation variables for parsing and calculations. Default would be latitude, longitude.
-gp (geophysical variables): Geophysical variables for parsing and calculations (e.g. aerosol optical depth land and ocean, solar azumith, etc).
-gs (gridding size): Gridding size and pixel resolution level.
-o (output): Output location
-on (output name): Output name. If there are multiple files created, output name will be the prefix and appended to time interval associated with the calculations.
-l (limit): Boundary box for latitude longitude (-90 90 180 180 would encompass the full Earth)
-r (read): Reads in raw data from L2 file.
-f (filter): Reads in raw data from L2 file and filters based on metadata.
-g (grid): Reads, filters, and grids single L2 file.
-ns (netCDF single): Reads, filters, grids single L2 file and saves output as netCDF file.
-nm (netCDF multiple): Reads, filters, grids single L2 file and saves output as single netCDF file regardless of time interval.
-nmt (netCDF multiple time): Reads, filters, grids single L2 file and saves output as single netCDF file with time interval separation as a layer dimension.
-ss (sensor statistics): Reads sensors and reports statistics and individual gridded data.
-sss (sensor statistic split): Reads sensors and reports statistics and gridded data based on satellite categorization.
-ssi (sensor statistic split id): Reads sensors and reports statistics and gridded data based on satellite categorization.
-cfg (config): Reads in YAML file and executes commands.
Command line input to call YAML file:
python3 gtools.py -cfg -fn "C:\LOCATION\CONFIG_FILE_NAME.yml"
The command line also has the ability to use YAML file while specifying the time start and end in the command line. This way there is no need to edit the YAML file every time when run (or create a new Docker image).
Command line input to call the YAML file with time start and end:
python3 gtools.py -cfgtime -fn "C:\LOCATION\CONFIG_FILE_NAME.yml" -ts 2020/01/01/00/00 -te 2020/01/01/00/30
The YAML file also has inputs for user specifications. This includes:
When reading a directory with subdirectories (e.g. LAADS archive), input path to the top directory in file_directory_folder. This would then read all files contained in subdirectories.
When creating a text file with paths to files, input path to this text file in file_directory_file. Paths should be included: C:\LOCATION\SATELLITE1.nc C:\LOCATION\SATELLITE2.hdf
The repository includes a Dockerfile, which was used to a build a Docker image for the package, which is available here: https://hub.docker.com/repository/docker/neilgutkin/aerosol-data-fusion/general.
A Docker image is essentially a blueprint for the creation of a Docker container. A container run from the image is a host-isolated environment that can be used to execute the data fusion package with provided user inputs.
Configuration of a YAML file is required for the package to be run with Docker. Through this configuration, the user specifies the various parameters for the package run. The template for this YAML file is available in the source directory of this repository, under the name "example_config.yml". The input, output, and static file location fields in the YAML should be set to the paths of the input and output as they appear in the container - the "file_io" section of the example config is already set up for the provided Docker image, so there is no need to change it.
The next step is setting up the file system on the host. The input file directory, output file directory, config.yml file, and static file must all be grouped into one directory on the host machine, referred to as the "ioFiles" directory in the example below.
Finally, it's time to run a container from the Docker image. This step requires the user to specify the location of the ioFiles directory that the package should use. This data will be shared between the container environment and the host, meaning that changes made in the container (e.g. by the package) will be reflected on the host. To run the container, a user can execute the following command:
docker run [flags] -v "/your/host/path/to/ioFilesDirectory:/app/src/ioFiles" [image_name]:[version] python ./gridtools/gtools.py -cfg -fn /app/src/ioFiles/config.yml
Below is an example - note especially the appearance of the windows source path (/c/ instead of C:/):
docker run -it --gpus all -v "/c/Users/Neil/Desktop/Work/s23/ioFiles:/app/src/ioFiles" aerosol-df:v0 python ./gridtools/gtools.py -cfg -fn /app/src/ioFiles/config.yml
If running on s4psci or a similar server environment, permissions might require that you add the :z flag to the command when linking directories. In the above example, you would replace "/c/Users/Neil/Desktop/Work/s23/ioFiles:/app/src/ioFiles" with "/c/Users/Neil/Desktop/Work/s23/ioFiles:/app/src/ioFiles:z". After execution, the package will run and the output files directory on the host machine will be populated with the newly fused outputs.
Navigate to the gridtools folder to run commands:
-$ python3 gtools.py [commands] [flags]
One example of this (run the yaml config file):
-$ python3 gtools.py -cfg -fn "PATH/config.yml"
where "-cfg" is the "config yaml" command and "-fn" is the filename flag for the proceeding path.
Inputs can be of the form of netCDF4 or HDf4 files. Sample files can be found in the respective folder in the "SampleInputs 0000-0059 01-01-2020" folder. These files can be found on NASA LAADS DAAC: https://ladsweb.modaps.eosdis.nasa.gov/archive/allData/
One can also request files at the NASA website (look under Atmosphere-Aerosol): https://ladsweb.modaps.eosdis.nasa.gov/search/
Outputs are in the form of netCDF4 file. Sample output files can be found in the respective folder in the "SampleOutputs 0000-0059 01-01-2020" folder.
Each output file is the fused statistics and grid for the input files for that time interval. If input files range between 00:00-23:59 for a single day and the time interval is 30 minutes, there will be 48 files produced (each of which is for that 30 minute time interval). These times can be changed by user preference.
The output files here use the sample input files provided and grid/fuse/provide statistics for Optical_Depth_Land_And_Ocean and Solar_Azumith between the times of 00:00 - 01:00, Jan 1 2020.
FAQs
Data fusion package for transforming L2 satellite to L3 spatial-temporal gridded data
We found that pyroscopegriddingcpu demonstrated a healthy version release cadence and project activity because the last version was released less than a year ago. It has 2 open source maintainers collaborating on the project.
Did you know?

Socket for GitHub automatically highlights issues in each pull request and monitors the health of all your open source dependencies. Discover the contents of your packages and block harmful activity before you install or update your dependencies.

Product
Ensure open-source compliance with Socket’s License Enforcement Beta. Set up your License Policy and secure your software!
Product
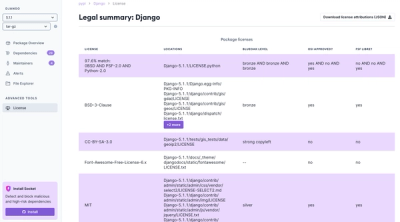
We're launching a new set of license analysis and compliance features for analyzing, managing, and complying with licenses across a range of supported languages and ecosystems.
Product
We're excited to introduce Socket Optimize, a powerful CLI command to secure open source dependencies with tested, optimized package overrides.