Product
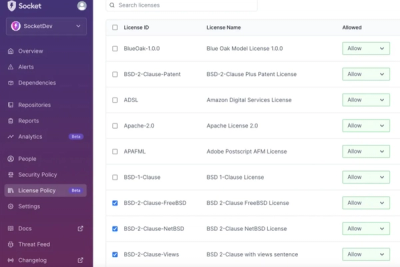
Introducing License Enforcement in Socket
Ensure open-source compliance with Socket’s License Enforcement Beta. Set up your License Policy and secure your software!
This is a library to evaluate an aminoacid sequence and determine an amphipathic index for each alpha helix or beta sheet.
This library can analyze an aminoacid sequence and gives a list of secondary structures with the respective hydrophobicity mean[1] and amphipathic index[1].
When it is useful to calculate this measurements on a secondary structure?
By looking into this measurements for an alpha helix, you can test some hiphotesis related with:
If you want to use this library on any GNU/Linux or OSX system you just need to execute:
$ pip install amphipathic
If you want to collaborate with this library, you should download the github repository and execute:
$ make deploy
To test all the project you should use the command:
$ pytest tests
This library can analyze an aminoacid sequence and gives a list of secondary structures with the respective hydrophobicity mean and amphipathic index.
import amphipathic
resume = amphipathic.index('NLYIQWLKDGGPSSGRPPPS')
print resume
or specifing scale="prift" from Cornette et al.[1]
import amphipathic
resume = amphipathic.index('NLYIQWLKDGGPSSGRPPPS', scale="prift")
print resume
And the result should be:
[[
{'end': 2,
'begin': 0,
'type': u'c',
'seq': 'nl',
'amphipathic': {'index': 7.572935321054872e-05, 'mean': 2.6}},
{'end': 5,
'begin': 2,
'type': u'e',
'seq': 'yiq',
'amphipathic': {'index': 1.4312912272216411, 'mean': 1.7299999999999998}},
{'end': 18,
'begin': 5,
'type': u'c',
'seq': 'wlkdggpssgrpp',
'amphipathic': {'index': 0.002511560979331271, 'mean': -0.43}},
{'end': 20,
'begin': 18,
'type': u'e',
'seq': 'ps',
'amphipathic': {'index': 1.6242872515167746, 'mean': -1.34}}
]]
Each secondary structure block has specific information like:
type could be "c" (from coil), "e" (extended/beta sheet) or "h" (alpha helix).mean provides the hydrophobicity mean obtained using the aminoacids of the block through Hydrophobicity scales obtained from Table 4 (STA PRIFT **) Cornette et al.[1].index provides an amphipathic index adapted from Cornette et al.[1], first implemented into Pablo Daniel Ghiringhelli's PhD thesis[2]. Cornette et al.[1] suggests an scalar equal or greater than 2, means apmhipathicity. On alpha helix cases this is only valid for segments shorter than 20-25 residues. When an alpha helix go further in longitude, the index is not valid anymore.It also accept a nucleotide sequence to perform the same analysis:
import amphipathic
resume = amphipathic.index('cgcgtccttggagcaatgcagttcaagaccaagaatcgaattgaacctgt')
print resume
And the output:
[[
{'end': 12,
'begin': 0,
'type': u'c',
'seq': 'rvlgamqfktkn',
'amphipathic': {'index': 0.007560225956225585, 'mean': 0.7825000000000001}},
{'end': 15,
'begin': 12,
'type': u'e',
'seq': 'rie',
'amphipathic': {'index': 1.6297837670649824, 'mean': 1.4599999999999997}},
{'end': 16,
'begin': 15,
'type': u'c',
'seq': 'p',
'amphipathic': {'index': 0.0, 'mean': -2.23}}
]]
Last, it also accept a polyprotein sequence. When working with aminoacid it detect the '*' character as a stop signal:
import amphipathic
resume = amphipathic.index('NLYIQWLKDG*GPSSGRPPPS')
print resume
This library was used into mistic2.
[1] Cornette, J. L., Cease, K. B., Margalit, H., Spouge, J. L., Berzofsky, J. A., & DeLisi, C. (1987). Hydrophobicity scales and computational techniques for detecting amphipathic structures in proteins. Journal of Molecular Biology, 195(3), 659–685. doi:10.1016/0022-2836(87)90189-6.
[2] Ghiringhelli D (2002). Virus Junín: Clonado molecular y análisis estructural y funcional del RNA S y sus productos génicos, Facultad de Ciencias Exactas, Universidad Nacional de La Plata.
If you want to develope with us or have questions about this library, please file an issue in this repository so some of the project managers can get back to you. Please check the existing (and closed) issues to make sure your issue hasn't already been addressed.
FAQs
This is a library to evaluate an aminoacid sequence and determine an amphipathic index for each alpha helix or beta sheet.
We found that amphipathic demonstrated a healthy version release cadence and project activity because the last version was released less than a year ago. It has 1 open source maintainer collaborating on the project.
Did you know?

Socket for GitHub automatically highlights issues in each pull request and monitors the health of all your open source dependencies. Discover the contents of your packages and block harmful activity before you install or update your dependencies.

Product
Ensure open-source compliance with Socket’s License Enforcement Beta. Set up your License Policy and secure your software!
Product
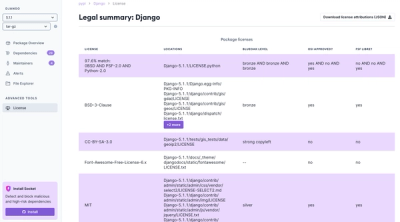
We're launching a new set of license analysis and compliance features for analyzing, managing, and complying with licenses across a range of supported languages and ecosystems.
Product
We're excited to introduce Socket Optimize, a powerful CLI command to secure open source dependencies with tested, optimized package overrides.