
Security News
Research
Data Theft Repackaged: A Case Study in Malicious Wrapper Packages on npm
The Socket Research Team breaks down a malicious wrapper package that uses obfuscation to harvest credentials and exfiltrate sensitive data.
Docs | Tutorials | Benchmarks | Papers Implemented
TorchDrug is a PyTorch-based machine learning toolbox designed for several purposes.
TorchDrug can be installed on either Linux, Windows or macOS. It is compatible with 3.7 <= Python <= 3.10 and PyTorch >= 1.8.0.
conda install torchdrug -c milagraph -c conda-forge -c pytorch -c pyg
pip install torch==1.9.0
pip install torch-scatter torch-cluster -f https://pytorch-geometric.com/whl/torch-1.9.0+cu102.html
pip install torchdrug
To install torch-scatter for other PyTorch or CUDA versions, please see the
instructions in https://github.com/rusty1s/pytorch_scatter
git clone https://github.com/DeepGraphLearning/torchdrug
cd torchdrug
pip install -r requirements.txt
python setup.py install
We need to first install the build tools for Visual Studio. We then install the following modules in PowerShell.
Install-Module Pscx -AllowClobber
Install-Module VSSetup
Initialize Visual Studio in PowerShell with the following commands. We may setup this for all PowerShell sessions by writing it to the PowerShell profile. Change the library path according to your own case.
Import-VisualStudioVars -Architecture x64
$env:LIB += ";C:\Program Files\Python37\libs"
We need PyTorch >= 1.13 to run TorchDrug on Apple silicon. For torch-scatter and
torch-cluster, they can be compiled from their sources. Note TorchDrug doesn't
support mps devices.
pip install torch==1.13.0
pip install git+https://github.com/rusty1s/pytorch_scatter.git
pip install git+https://github.com/rusty1s/pytorch_cluster.git
pip install torchdrug
TorchDrug is designed for humans and focused on graph structured data. It enables easy implementation of graph operations in machine learning models. All the operations in TorchDrug are backed by PyTorch framework, and support GPU acceleration and auto differentiation.
from torchdrug import data
edge_list = [[0, 1], [1, 2], [2, 3], [3, 4], [4, 5], [5, 0]]
graph = data.Graph(edge_list, num_node=6)
graph = graph.cuda()
# the subgraph induced by nodes 2, 3 & 4
subgraph = graph.subgraph([2, 3, 4])
Molecules are also supported in TorchDrug. You can get the desired molecule properties without any domain knowledge.
mol = data.Molecule.from_smiles("CCOC(=O)N", atom_feature="default", bond_feature="default")
print(mol.node_feature)
print(mol.atom_type)
print(mol.to_scaffold())
You may also register custom node, edge or graph attributes. They will be automatically processed during indexing operations.
with mol.edge():
mol.is_CC_bond = (mol.edge_list[:, :2] == td.CARBON).all(dim=-1)
sub_mol = mol.subgraph(mol.atom_type != td.NITROGEN)
print(sub_mol.is_CC_bond)
TorchDrug provides a wide range of common datasets and building blocks for drug discovery. With minimal code, you can apply standard models to solve your own problem.
import torch
from torchdrug import datasets
dataset = datasets.Tox21()
dataset[0].visualize()
lengths = [int(0.8 * len(dataset)), int(0.1 * len(dataset))]
lengths += [len(dataset) - sum(lengths)]
train_set, valid_set, test_set = torch.utils.data.random_split(dataset, lengths)
from torchdrug import models, tasks
model = models.GIN(dataset.node_feature_dim, hidden_dims=[256, 256, 256, 256])
task = tasks.PropertyPrediction(model, task=dataset.tasks)
Training and inference are accelerated by multiple CPUs or GPUs. This can be seamlessly switched in TorchDrug by just a line of code.
from torchdrug import core
# Single CPU / Multiple CPUs / Distributed CPUs
solver = core.Engine(task, train_set, valid_set, test_set, optimizer)
# Single GPU
solver = core.Engine(task, train_set, valid_set, test_set, optimizer, gpus=[0])
# Multiple GPUs
solver = core.Engine(task, train_set, valid_set, test_set, optimizer, gpus=[0, 1, 2, 3])
# Distributed GPUs
solver = core.Engine(task, train_set, valid_set, test_set, optimizer, gpus=[0, 1, 2, 3, 0, 1, 2, 3])
Experiments can be easily tracked and managed through Weights & Biases platform.
solver = core.Engine(task, train_set, valid_set, test_set, optimizer, logger="wandb")
Everyone is welcome to contribute to the development of TorchDrug. Please refer to contributing guidelines for more details.
TorchDrug is released under Apache-2.0 License.
FAQs
A powerful and flexible machine learning platform for drug discovery
We found that torchdrug demonstrated a healthy version release cadence and project activity because the last version was released less than a year ago. It has 1 open source maintainer collaborating on the project.
Did you know?

Socket for GitHub automatically highlights issues in each pull request and monitors the health of all your open source dependencies. Discover the contents of your packages and block harmful activity before you install or update your dependencies.

Security News
Research
The Socket Research Team breaks down a malicious wrapper package that uses obfuscation to harvest credentials and exfiltrate sensitive data.
Research
Security News
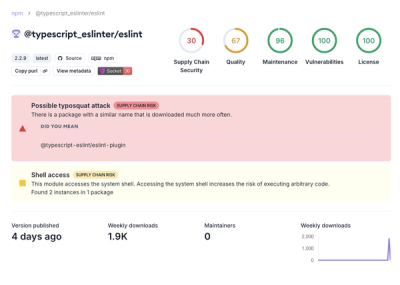
Attackers used a malicious npm package typosquatting a popular ESLint plugin to steal sensitive data, execute commands, and exploit developer systems.

Security News
The Ultralytics' PyPI Package was compromised four times in one weekend through GitHub Actions cache poisoning and failure to rotate previously compromised API tokens.